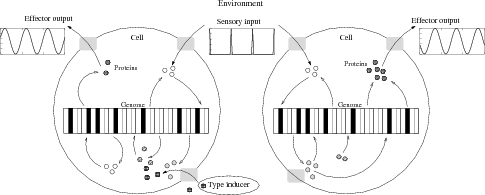
Schematic Protein / Genome / Environment interaction of differentiated cells.
Here you can find:
| Supporting and supplemental information for the paper "The Essential Motif That Wasn't There: Topological and Lesioning Analysis of Evolved Genetic Regulatory Networks" |

Schematic Protein / Genome / Environment interaction of differentiated cells. Here you can find: |
| What is in the paper? |
|
Abstract Networks that model natural Genetic Regulatory Networks (GRNs) are evolved to show a range of dynamical behaviors. Specifically one group was evolved to show differentiation, i.e. to be able to perform an additional behavior as compared to the original, single target function. These GRNs are then analyzed and compared with measures used in the biological sciences. Having huge numbers of GRNs available for not only analysis but also ``metabolic" inspection, we find that evolutionary niches (target functions) do not necessarily mold network structure uniquely.
Our results suggest that variability operators can have a stronger influence on network topologies than selection pressures, especially when many topologies can create similar dynamics.
Furthermore, damaging the most significantly represented motif (whether in differentiating or non-differentiating GRNs) is found not to have a bigger impact on function than random lesions, suggesting that particular motifs are not as important in the robust functioning of networks as might perhaps be expected. Full text available at IEEE Xplore |
| Network dynamics visualization with a Java Applet |
|
Please choose between three networks by selecting from the drop-down menu:
The source code for this applet can be found in the section "Java code". |
| Java source code |
Please feel free to download the full Java source code. Structuring of the code in quite a few classes has hopefully increased readability.
|
| Lookup tables mentioned in the paper |
Due to space reasons we could only mention that the four bit decay rate per protein,
the three bit saturation values and the four bit binding proportion parts of the genome
are decoded by taking values from a lookup table. Here you find the arrays from our
Java implementation (see code) used for generating the results reported in the paper:
public static double[] decayRates={0.0,0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0,1.0,1.0,1.0,1.0,1.0,1.0}; //\in [0..1]
public static double[] saturationValues={10,25,50,100,150,200,300,500};
public static double[] bindingProportions={0.0,0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0,1.0,1.0,1.0,1.0,1.0,1.0}; //\in [0..1]
|
| Additional results |
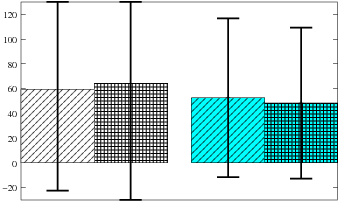
Impact of lesions on 4-node subgraphs: Average reduction in fitness when one edge in a network is removed for the original (left) and differentiated GRNs (right). In each pair, the diagonally striped bar shows the impact when any edge can be chosen while for the grid patterned bar an edge from the nets most significant network motif was removed. On average the differences are very small and the standard deviations huge, as in the results for 3-node subgraphs shown in the paper. 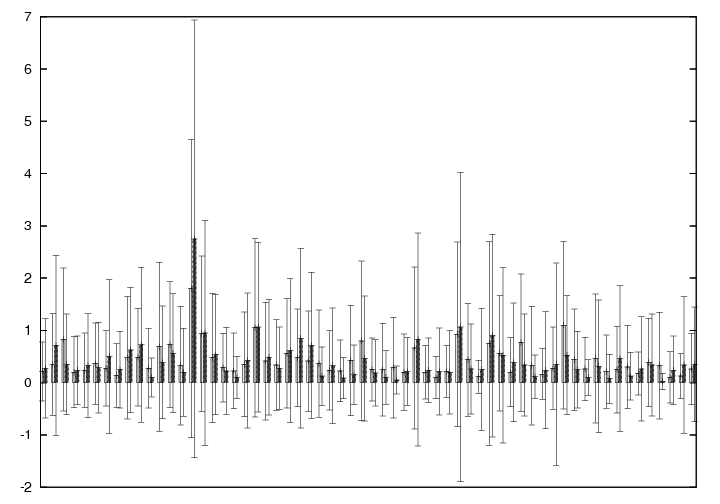
Evolved for differentiation (light gray) vs. evolved in the original setting for one task (dark gray) average network pattern distributions. Subgraph patterns are shown on the x-axis and average number of occurrences on the y-axis. Corresponds to figure 7 of the paper, only here results are for 4-node subgraphs - again we find average differences to be not very big.
|
| 3 node subgraphs |
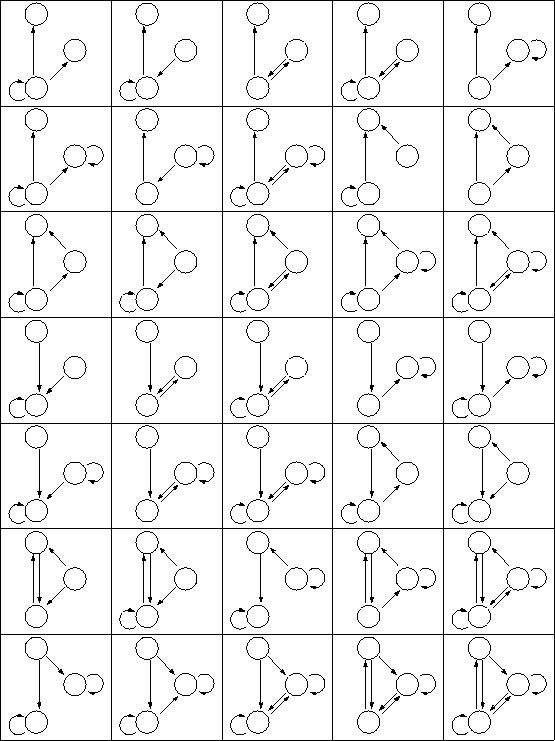
Reproduction from the paper: 3-node subgraph network patterns occuring more often than 0.2 times within evolved GRN populations on average. (There are too many 4-node subgraphs for reasonable cut-off values to show them here.) |
| Reference / Bibtex |
Knabe, J. F., Nehaniv, C. L. and Schilstra, M. J. The Essential Motif that wasn't there: Topological and Lesioning Analysis of Evolved Genetic Regulatory Networks. In IEEE Symposium on Artificial Life (CI-ALife'07), pages 69-76, Omnipress, 2007.
@InProceedings{GRNmotifs,
author = {Johannes F. Knabe and Chrystopher L. Nehaniv and Maria J. Schilstra},
title = {The Essential Motif that wasn't there: Topological and Lesioning Analysis of Evolved Genetic Regulatory Networks},
booktitle = {IEEE Symposium on Artificial Life (CI-ALife'07)},
note = {CI-ALife'07 was part of the 2007 IEEE Symposium Series on Computational Intelligence},
publisher = {Omnipress},
year = {2007},
pages = {69--76},
ee= {http://ieeexplore.ieee.org/iel5/4218857/4218858/04218870.pdf?tp=&arnumber=4218870&isnumber=4218858},
url = {http://panmental.de/GRNmotifs}
}
|