List of contents for jumping directly to a subsection:
- Motif distributions for nets restricted to 3, 4 or 5 gene nodes
- Animation showing the changing motif distribution over evolutionary time for nets contstrained to 5 gene nodes
The following results are all for 9 node networks:
- Table of all 4-node subgraph patterns found in significant numbers
- Impact of lesioning; random damage vs most significant 3-node motif damaged after 500 generations and random damage vs most significant 4-node motif damaged after 1000 generations
- Distribution of 3-node motif patterns after generations 500, 750 and 1000
- A lot of data on motif sub-structure distribution:
- Motif #7, sub-structure distribution after generation 1000 (not present above cut-off value before)
- Motif #13, sub-structure distribution after generations 500, 750 and 1000
- Motif #22, sub-structure distribution after generations 500, 750 and 1000
- Motif #31, sub-structure distribution after generation 1000 (not present above cut-off value before)
- Motif #37, sub-structure distribution after generations 500, 750 and 1000
- Motif #38, sub-structure distribution after generations 500, 750 and 1000
- Motif #45, sub-structure distribution after generations 500, 750 and 1000
- Motif #73, sub-structure distribution after generations 500, 750 and 1000
- Motif #74, sub-structure distribution after generations 500, 750 and 1000
- Motif #75, sub-structure distribution after generations 500, 750 and 1000
- Motif #82, sub-structure distribution after generations 500, 750 and 1000
- Motif #89, sub-structure distribution after generations 500, 750 and 1000
- Motif #90, sub-structure distribution after generations 500, 750 and 1000
- Motif #91, sub-structure distribution after generations 500, 750 and 1000
- Motif #105, sub-structure distribution after generations 500, 750 and 1000
- Motif #109, sub-structure distribution after generations 500, 750 and 1000
- Motif #211, sub-structure distribution after generations 500, 750 and 1000
- Motif #219, sub-structure distribution after generations 500, 750 and 1000
Results:
- Motif distributions for nets restricted to 3, 4 or 5 gene nodes
- Animation showing the changing motif distribution over evolutionary time for nets contstrained to 5 gene nodes
- Table of all 4-node subgraph patterns found in significant numbers
- Impact of lesioning; random damage vs most significant 3-node motif damaged after 500 generations and random damage vs most significant 4-node motif damaged after 1000 generations
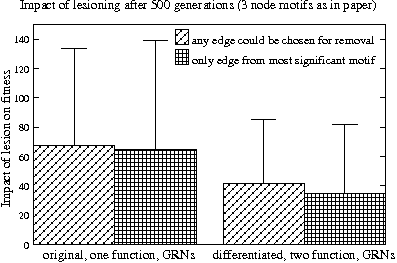
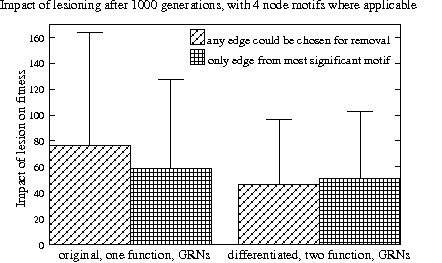 Average reduction in fitness when one edge in a network is removed for the original (left) and differentiated GRNs (right). In each pair, the diagonally striped bar shows the impact when any edge can be chosen while for the grid patterned bar an edge from the nets most significant network motif was removed. On average the differences are very small and the standard deviations huge.
Average reduction in fitness when one edge in a network is removed for the original (left) and differentiated GRNs (right). In each pair, the diagonally striped bar shows the impact when any edge can be chosen while for the grid patterned bar an edge from the nets most significant network motif was removed. On average the differences are very small and the standard deviations huge.
go back to top of additional results section
- Distribution of 3-node motif patterns after generations 500, 750 and 1000
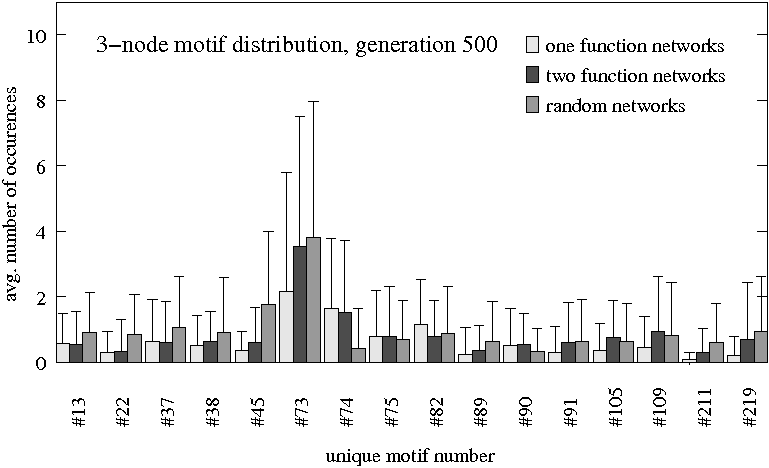
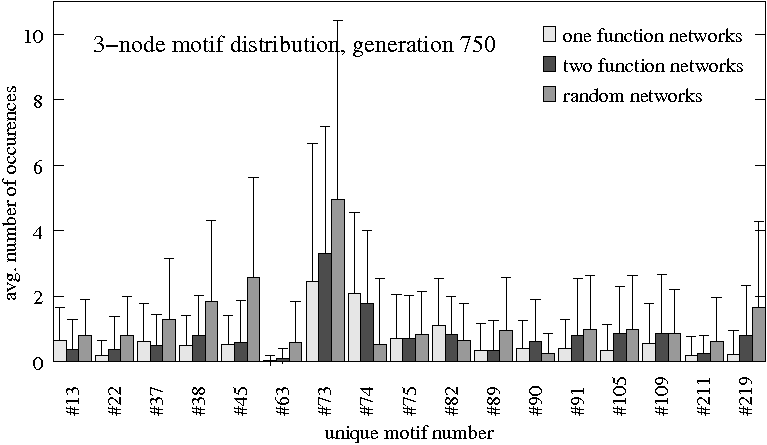
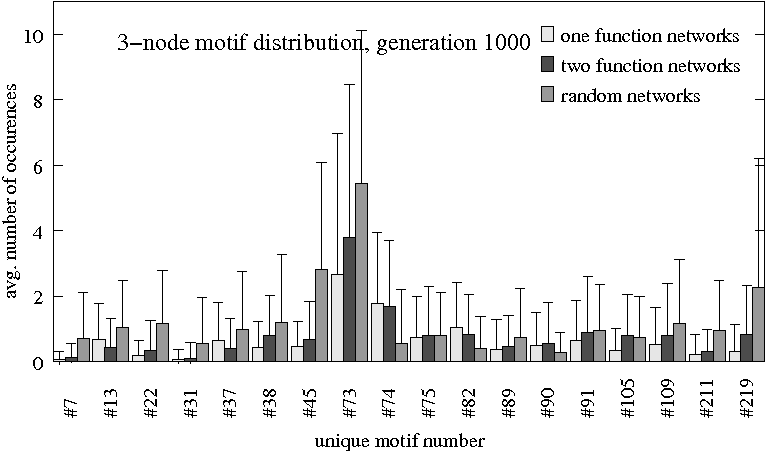 Three groups ($N=80$ each) are compared: Original evolutionary setting for one task (light gray), evolved for differentiation (dark gray) and randomly evolved (medium gray). Only subgraph patterns that have more than the cut-off value of 0.5 occurences on average in at least one of the groups are shown.
Three groups ($N=80$ each) are compared: Original evolutionary setting for one task (light gray), evolved for differentiation (dark gray) and randomly evolved (medium gray). Only subgraph patterns that have more than the cut-off value of 0.5 occurences on average in at least one of the groups are shown.
go back to top of additional results section
- Motif #7, sub-structure distribution after generation 1000 (not present above cut-off value before)
- Motif #13, sub-structure distribution after generations 500, 750 and 1000
- Motif #22, sub-structure distribution after generations 500, 750 and 1000
- Motif #31, sub-structure distribution after generation 1000 (not present above cut-off value before)
- Motif #37, sub-structure distribution after generations 500, 750 and 1000
- Motif #38, sub-structure distribution after generations 500, 750 and 1000
- Motif #45, sub-structure distribution after generations 500, 750 and 1000
- Motif #73, sub-structure distribution after generations 500, 750 and 1000
- Motif #74, sub-structure distribution after generations 500, 750 and 1000
- Motif #75, sub-structure distribution after generations 500, 750 and 1000
- Motif #82, sub-structure distribution after generations 500, 750 and 1000
- Motif #89, sub-structure distribution after generations 500, 750 and 1000
- Motif #90, sub-structure distribution after generations 500, 750 and 1000
- Motif #91, sub-structure distribution after generations 500, 750 and 1000
- Motif #105, sub-structure distribution after generations 500, 750 and 1000
- Motif #109, sub-structure distribution after generations 500, 750 and 1000
- Motif #211, sub-structure distribution after generations 500, 750 and 1000
- Motif #219, sub-structure distribution after generations 500, 750 and 1000
|